Older Projects
This is a list of research projects that we have undertaken in the past that are no longer active. We also provide here resources that were produced in these projects and are freely available, but which are no longer supported.
Metabolic Reconstructions
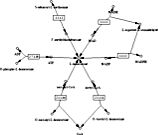

This was a collaboration with many groups around the world. A basic step in understanding cellular biology is to know all of its components and how they interact. A metabolic reconstruction is a network of molecules, reactions and genes involved in cellular metabolism. We have been involved in several metabolic reconstructions, but particularly for yeast and human (Recon2).
- Swainston N, Smallbone K, Hefzi H, Dobson PD, Brewer J, Hanscho M, Zielinski DC, Siong Ang K, Gardiner NJ, Gutierrez JM, Kyriakopoulos S, Lakshmanan M, Li S, Liu JK, Martínez VS, Orellana CA, Quek L-E, Thomas A, Zanghellini J, Borth, N, Lee D-Y, Nielsen LK, Kell DB, Lewis NE, Mendes P (2016) Recon 2.2: from reconstruction to model of human metabolism. Metabolomics 12:109 [full text]
- Thiele I, Swainston N, Fleming RMT, Hoppe A, Sahoo S, Aurich MK, Haraldsdottir H, Mo ML, Rolfsson O, Stobbe MD, Thorleifsson SG, Agren R, Bölling C, Bordel S, Chavali AK, Dobson P, Dunn WD, Endler L, Hala D, Hucka M, Hull D, Jameson D, Jamshidi N, Jonsson JJ, Juty N, Keating S, Nookaew I, Le Novère N, Malys N, Mazein A, Papin JA, Price ND, Selkov Sr E, Sigurdsson MI, Simeonidis E, Sonnenschein N, Smallbone K, Sorokin A, van Beek JHGM, Weichart D, Goryanin I, Nielsen J, Westerhoff HV, Kell DB, Mendes P & Palsson BØ (2013) A community-driven global reconstruction of human metabolism Nature Biotechnology 31:419-425 [full text]
- Herrgård MJ, Swainston N, Dobson P, Dunn WB, Arga KY, Arvas M, Blüthgen N, Borger S, Costenoble R, Heinemann M, Hucka M, Le Novère N, Li P, Liebermeister W, Mo ML, Oliveira AP, Petranovic D, Pettifer S, Simeonidis E, Smallbone K, Spasić I, Weichart D, Brent R, Broomhead DS, Westerhoff HV, Kirdar B, Penttilä M, Klipp E, Palsson BØ, Sauer U, Oliver SG, Mendes P, Nielsen J, Kell DB (2008) A consensus yeast metabolic network reconstruction obtained from a community approach to systems biology, Nature Biotechnol. 26, 1155-1160. [abstract] [full text].
Mathematical modeling of yeast oxidative stress

This was a collaboration with R. Laubenbacher and V. Shulaev at the Virginia Bioinformatics Institute funded by the NIGMS and NSF. We conducted a systems biology study of the dynamics of the response of yeast to cumene hydroperoxyde through microrarrays, proteomics and metabolomics.
- Sha W, Martins AM, Laubenbacher R, Mendes P, Shulaev V (2013) The genome-wide early temporal response of Saccharomyces cerevisiae to oxidative stress induced by cumene hydroperoxide. PLOS One 8: e74939. [full text] [supplementary data]
- Martins, A.M., Sha, W., Evans, C., Martino-Catt, S., Mendes, P., Shulaev, V. (2007) Comparison of sampling techniques for parallel analysis of transcript and metabolite levels in Saccharomyces cerevisiae. Yeast 24, 181-188. [abstract]
- Martins, A.M., Camacho, D., Shuman, J., Sha, W., Mendes, P. & Shulaev, V. (2004) A systems biology study of two distinct growth phases of Saccharomyces cerevisiae. Current Genomics 5, 649-663. [abstract]
Metabolic Engineering of Plant Vitamin C Biosynthesis

This was a collaborative project with Craig Nessler, Argelia Lorence, and Boris Chevone at Virginia Tech funded by NSF and USDA. We discovered new myo-inositol oxigenase enzymes in Arabidopsis. Overexpressing these genes leads to plants with larger biomass (particularly roots).
- Lisko KA, Torres R, Harris RS, Belisle M, Vaughan MM, Jullian B, Chevone BI, Mendes P, Nessler CL, Lorence A (2013) Elevating vitamin C content via over-expression of myo-inositol oxygenase and L-gulono-1,4-lactone oxidase in Arabidopsis leads to enhanced biomass and tolerance to abiotic stresses. In Vitro Cellular and Developmental Biology - Plant 49:643-655. [abstract]
- Lorence, A., Chevone, B.I., Mendes, P. & Nessler, C.L. (2004) myo-Inositol oxygenase offers a possible entry point into plant ascorbate biosynthesis. Plant Physiology 134, 1200-1205. [abstract] [full text]
Medicago truncatula functional genomics and bioinformatics

This was a collaboration with Rick Dixon of the S.R. Noble Foundation and was funded by the National Science Foundation. More details about this project on its NSF Plant Genome Research Program.
- Lei, Z., Elmer, A.M., Watson, B.S., Dixon, R.A., Mendes, P., Sumner, L.W. (2005) A two-dimensional electrophoresis proteomic reference map and systematic identification of 1367 proteins from a cell suspension culture of the model legume Medicago truncatula. Mol. Cell. Proteomics 4 1812-1825 [abstract] [full text]
- Suzuki, H., Reddy, M.S., Naoumkina, M., Aziz, N., May, G.D., Huhman, D.V., Sumner, L.W., Blount, J.W., Mendes, P. & Dixon, R.A. (2005) Methyl jasmonate and yeast elicitor induce differential transcriptional and metabolic re-programming in cell suspension cultures of the model legume Medicago truncatula. Planta 220, 696-707 [abstract] [full text]
- Broeckling, C.D., Huhman, D.V., Farag, M.A., Smith, J.T., May, G.D., Mendes, P., Dixon, R.A. & Sumner, L.W. (2005) Metabolic profiling of Medicago truncatula cell cultures reveals the effects of biotic and abiotic elicitors on metabolism. Journal of Experimental Botany 56, 323-336. [abstract] [full text]
- Sumner, L.W., Mendes, P., Dixon, R.A. (2003) Plant metabolomics: large-scale phytochemistry in the functional genomics era. Phytochemistry 62, 817-36. [abstract] [full text]
Grape Bioinformatics

This was a collaborative project led by Grant Cramer from the University of Nevada-Reno, and funded by the NSF Plant Genome Research Program. More information at the project's main website.
- Cramer, G.R., Cushman, J.C., Schooley, D.A., Quilici, D., Vincent, D., Bohlman, M.C., Ergul, A., Tattersall, E.A.R., Tillett, R., Evans, J., Delacruz, R., Schlauch, K. & Mendes, P. (2005) Progress in bioinformatics - the challenge of integrating transcriptomic, proteomic and metabolomic information. Acta Hort. 689, 417-425 [abstract]
DOME
The DOME database system was constructed to support the data obtained from the Medicago, grape and yeast projects above. This system is no longer being developed, however we are happy to distribute the software to anyone who wants to use it. Unfortunately there are no means for us to provide support.
- Mehrotra, B., Mendes, P. (2006) Bioinformatics approaches to integrate metabolomics and other systems biology data. in Plant Metabolomics (Biotechnology in Agriculture and Forestry) (Saito, K., Dixon, R.A. & Willmitzer, L. Eds.) Springer-Verlag, Berlin and Heidelberg, Germany, pp. 105-116.
- Li, X.J., Brazhnik, O., Kamal, A., Guo, D., Lee, C., Hoops, S. & Mendes, P. (2002) Databases and visualization for metabolomics. In Harrigan, G.G. and Goodacre, R. (eds.), Metabolic profiling: its role in biomarker discovery and gene function analysis Kluwer Academic Publishers, Boston, Dordrecht and London, pp. 293-309.
